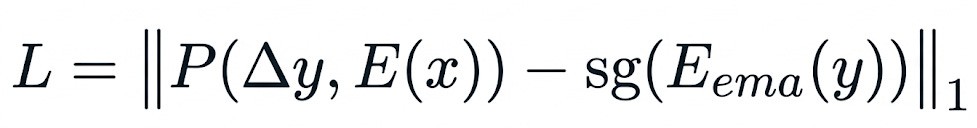EchoJEPA: How We Trained AI on 18 Million Heart Videos - With Zero Labels
Introducing EchoJEPA: the world's best video foundation model for echocardiography.
What if I told you an AI model could watch millions of heart ultrasound videos - with zero human annotations - and learn, on its own, how the heart moves, which chambers it’s looking at, and how well the heart is pumping?
No labels. No expensive cardiologist hours spent painstakingly annotating frames. Just raw video. Millions of hours of hearts beating.
And then, when you finally give it just 1% of labeled data, it destroys every existing method on the planet.
That’s EchoJEPA. And I’m thrilled to share that I co-authored this work.
📄 Read the Paper | 🌐 Project Page | 💻 Code
Why Echocardiography AI Needs a New Approach?
Echocardiography - heart ultrasound - is the most commonly used cardiac imaging modality in the world. Around 30 million echo studies are performed annually in the United States alone. It’s how doctors see your heart pump in real time. No radiation. No invasive procedures. Just a probe, some gel, and sound waves bouncing off your heart.
But here’s the problem: training AI models on echo data has historically required massive amounts of expert-labeled data. Think cardiologists manually clicking through thousands of videos to mark heart chambers, trace walls, measure how well the heart squeezes. That process is expensive, slow, and simply doesn’t scale.
Now multiply that by the diversity of hospitals, ultrasound machines, patient populations, body types, and image quality variations across the globe.
The result? Most echo AI models work well in the hospital where they were trained - and fall apart everywhere else.
EchoJEPA changes this equation entirely.
The Big Idea (In Plain English)
Imagine teaching a child about animals.
Approach A: Show them 100 flashcards with labels - “this is a dog,” “this is a cat.” That’s supervised learning. It works, but it’s slow, and the child only recognizes the animals you labeled.
Approach B: Let the child watch thousands of hours of nature documentaries. No labels. Just observation. The child starts noticing patterns on their own - things that move a certain way, things with similar shapes, things that appear together. When you finally show them one flashcard and say “this is a dog,” they get it instantly. They have already built an internal model of the world.
That’s what EchoJEPA does. Except the “nature documentaries” are 18 million echocardiography videos from over 300,000 patients - the largest echo pretraining dataset ever assembled. And instead of memorizing pixels, it learns abstract representations of cardiac anatomy and motion.
The model watches hearts beat. It learns what matters. And when you finally give it a tiny bit of guidance, it outperforms everything else - by a lot.
Why Existing Methods Fall Short
Before EchoJEPA, the field had three main approaches to building echo AI foundation models. Each has a fundamental limitation:
Supervised multitask learning (like PanEcho): Train on labeled data across many tasks simultaneously. The problem? You inherit all the noise and inconsistency in your annotations. And you need labels - lots of them.
Contrastive vision-language learning (like EchoPrime): Align video representations with text from clinical reports. Clever, but the model learns to match report language rather than cardiac anatomy. A report that says “normal LV function” doesn’t teach the model what normal LV function looks like at the tissue level.
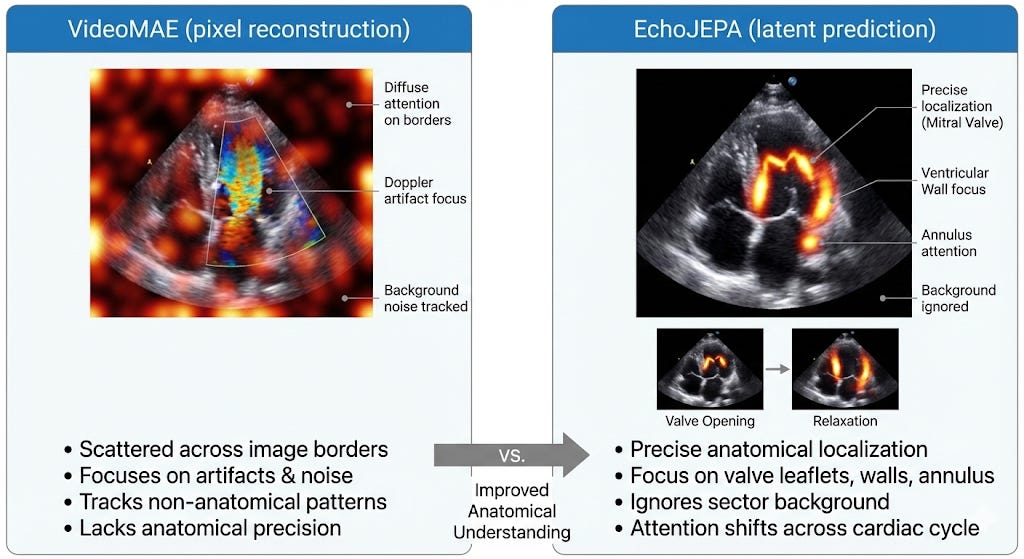
Masked pixel reconstruction (like EchoFM and VideoMAE): Hide parts of the video and train the model to fill in the missing pixels. This is where it gets interesting - and where EchoJEPA’s key insight comes in.
Here’s the critical problem with pixel reconstruction in ultrasound: echo images are dominated by speckle noise. Speckle is that grainy, textured pattern you see in every ultrasound image. It’s an inherent property of ultrasound physics - it comes from the constructive and destructive interference of scattered sound waves from tissue microstructure. It varies randomly between acquisitions. And it has nothing to do with whether the heart is healthy or sick.
A pixel-reconstruction model doesn’t know that. It will waste enormous capacity faithfully learning to reproduce noise patterns that carry zero diagnostic information. It’s like asking a student to memorize the exact grain pattern of a photograph instead of understanding what the photograph depicts.
EchoJEPA takes a fundamentally different path.
How EchoJEPA Works (The Technical Core)
EchoJEPA is built on Meta’s V-JEPA (Video Joint Embedding Predictive Architecture), adapted and scaled specifically for cardiac ultrasound. The core idea, originally proposed by Yann LeCun, is deceptively simple but profoundly effective.
Latent Prediction, Not Pixel Reconstruction
Instead of predicting missing pixels, EchoJEPA predicts missing representations in an abstract latent space.
Here’s how it works:
Partition and mask. The input video is divided into spatio-temporal “tubelets” - small 3D patches spanning both space and time. A large portion of these tubelets are masked (hidden from the model).
Encode the visible context. A context encoder (a Vision Transformer) processes only the visible, unmasked tubelets and produces embeddings - compact mathematical representations of what it sees.
Predict the missing parts - in latent space. A predictor network takes the visible embeddings plus positional information about where the masked tubelets are, and tries to infer what the embeddings of those masked regions should be. Not the pixels. The meaning.
Compare against a stable target. The prediction is compared against targets produced by an Exponential Moving Average (EMA) encoder - a slowly-updating copy of the main encoder. This is trained with an L₁ loss. The EMA target provides stable learning signals and, critically, naturally suppresses stochastic noise like speckle.
Formally, the loss function is:
where P is the predictor, E is the context encoder, E_ema is the EMA target encoder, x are the visible tubelets, y are the masked tubelets, and sg(·) is the stop-gradient operator that prevents representational collapse.
Why does this matter for ultrasound? Because the EMA target encoder evolves slowly, it captures only what is consistently present across training - anatomical structures, chamber geometry, wall motion. The random speckle, which changes unpredictably between frames and acquisitions, gets naturally filtered out. The model learns to represent what matters and ignore what doesn’t.
This is not a small implementation detail. This is the entire thesis of our paper. And the results prove it decisively.
Domain Adaptations: Respecting Cardiac Physics
We didn’t just take V-JEPA off the shelf and throw echo data at it. We made three targeted adaptations:
Higher temporal resolution (4 fps → 24 fps). Cardiac events happen fast - a mitral valve opens and closes in 50–100 milliseconds. Standard video frame rates miss critical dynamics. We needed the model to see the heart’s rapid motion.
Narrower aspect ratio augmentation (0.75–1.35 → 0.9–1.1). Echocardiographic views follow standardized acquisition protocols. Aggressive aspect ratio distortion warps chamber proportions and destroys clinically meaningful geometry. We kept augmentation tight to preserve what cardiologists actually measure.
Conservative crop scale (0.3–1.0 → 0.5–1.0). Ultrasound images have a characteristic fan-shaped sector. Crop too aggressively and you lose the heart entirely. We ensured every crop retains meaningful cardiac structures.
These may sound like minor engineering choices. They’re not. They’re the difference between a model that understands cardiac anatomy and one that hallucinates.
Scale: 1.1 Billion Parameters, 18 Million Videos
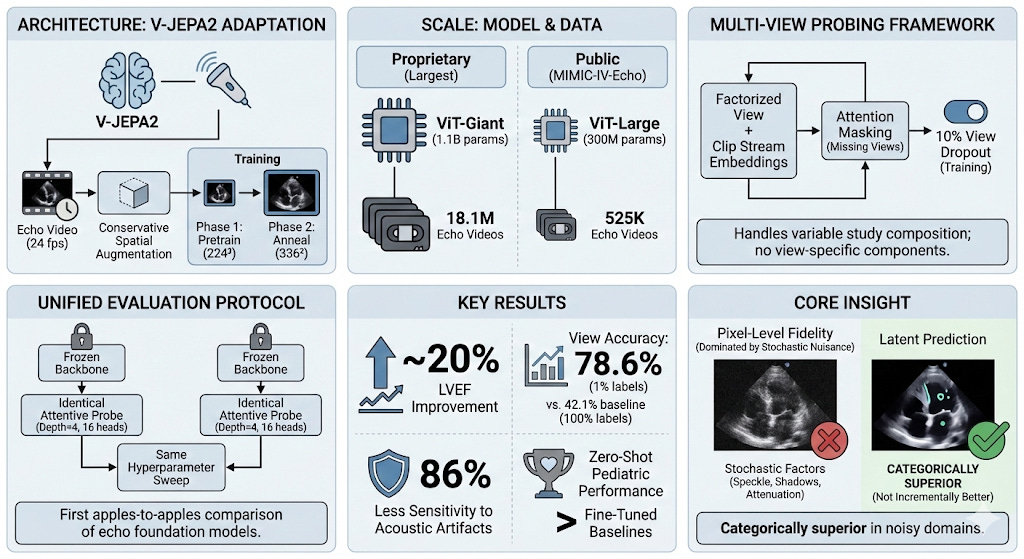
Our flagship model, EchoJEPA-G, uses a ViT-Giant backbone with 1.1 billion parameters, pretrained on 18.1 million proprietary echocardiography videos spanning diverse patient populations, hospitals, and scanner manufacturers. Training proceeded in two phases: pretraining at 224² resolution for 280 epochs, then annealing at 336² for 80 epochs.
This is the largest echocardiography pretraining effort ever undertaken.
For reproducibility, we also release EchoJEPA-L (ViT-Large, 300M parameters) trained on 525K videos from the public MIMIC-IV-Echo dataset - so anyone can validate our methodology.
Multi-View Probing: Seeing the Full Picture
A real echo study isn’t a single video — it’s a collection of clips from multiple views (apical 4-chamber, parasternal long axis, subcostal, etc.). Different clinical questions require different combinations of views. For instance, estimating right ventricular systolic pressure (RVSP) requires integrating information from apical views (to measure tricuspid regurgitation velocity) and subcostal views (to assess IVC diameter).
We developed a multi-view attentive probing framework that handles this naturally. Each view’s video is encoded independently by the frozen EchoJEPA backbone. The resulting embeddings are augmented with learnable view and clip stream embeddings (factorized, so we need only (V + C) × D parameters instead of V × C × D), concatenated, and passed through a lightweight cross-attention probe.
A key innovation: 10% view dropout during training randomly masks entire views, forcing the model to make valid predictions even with incomplete studies. This directly addresses the clinical reality that not every patient gets a complete, textbook-quality echo study.
All of this - and I want to emphasize this - operates on top of a completely frozen backbone. The encoder weights are never updated. Only the lightweight probe is trained. This means differences in performance reflect pure representation quality, not fine-tuning tricks.
The Results: State-of-the-Art Across the Board
Let me walk you through what happened when we put EchoJEPA against the best existing methods. All comparisons use frozen backbones with identical lightweight probes - no architectural advantages. Pure representation quality, head to head.
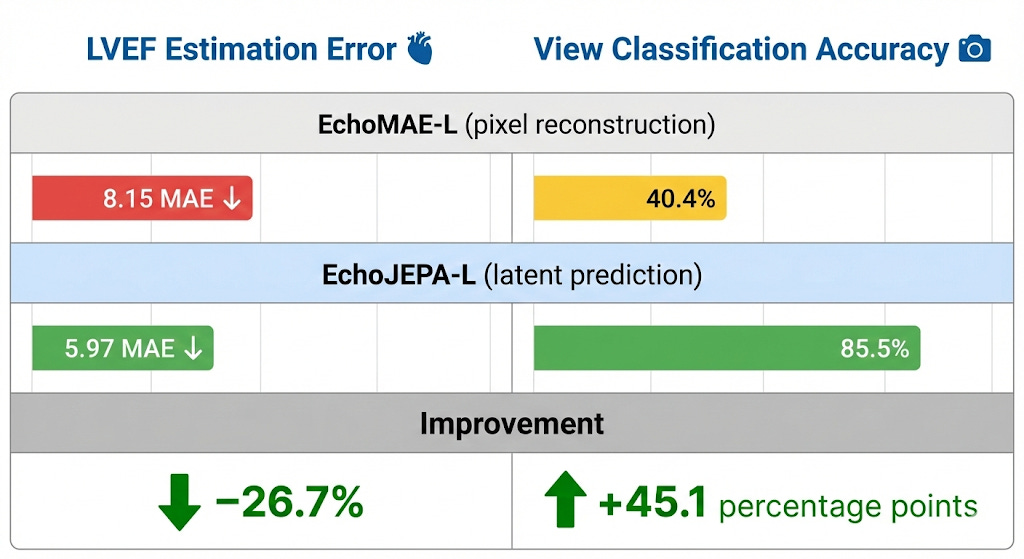
1. Latent vs. Pixel Prediction (The Controlled Experiment)
This is the cleanest experiment in the paper. Same ViT-Large architecture. Same MIMIC-IV-Echo data (525K videos). Same augmentations. Same compute budget on 8×H100 GPUs. The only difference: latent prediction (EchoJEPA-L) vs. pixel reconstruction (EchoMAE-L).
Same data. Same compute. Radically different objective. Radically different results. Latent prediction isn’t marginally better for ultrasound - it’s a paradigm shift.
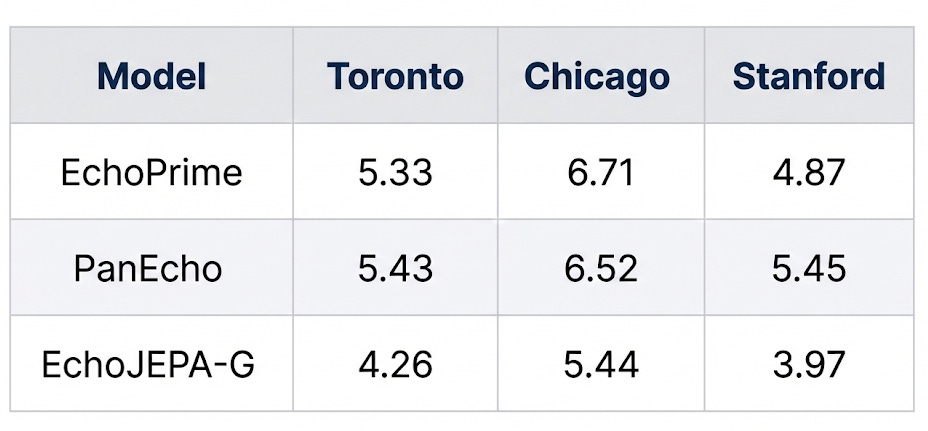
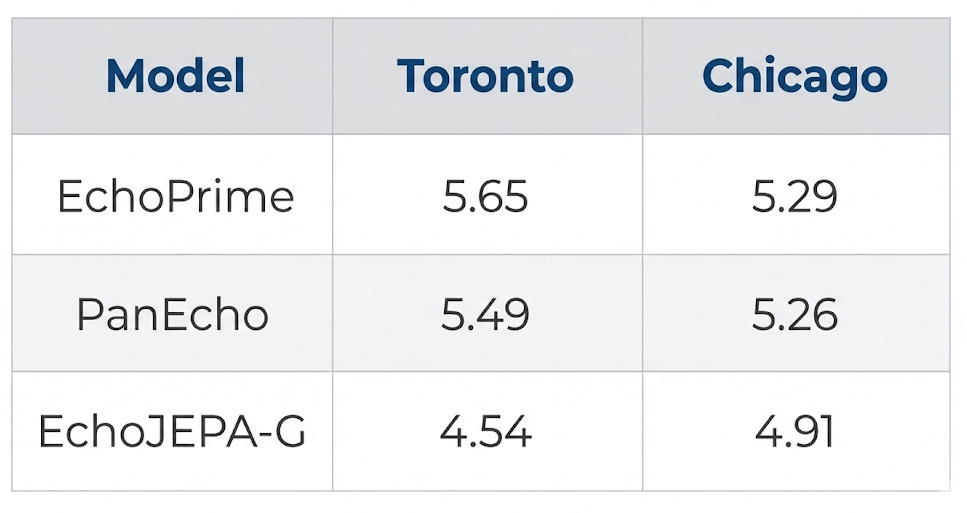
2. State-of-the-Art LVEF Estimation
Left ventricular ejection fraction (LVEF) is the single most important number in cardiology - it tells you how well the heart pumps blood. Lower MAE = better.
EchoJEPA-G achieves the best performance at every single site - including sites it was never trained on. A ~20% improvement over the previous best on the Toronto benchmark.
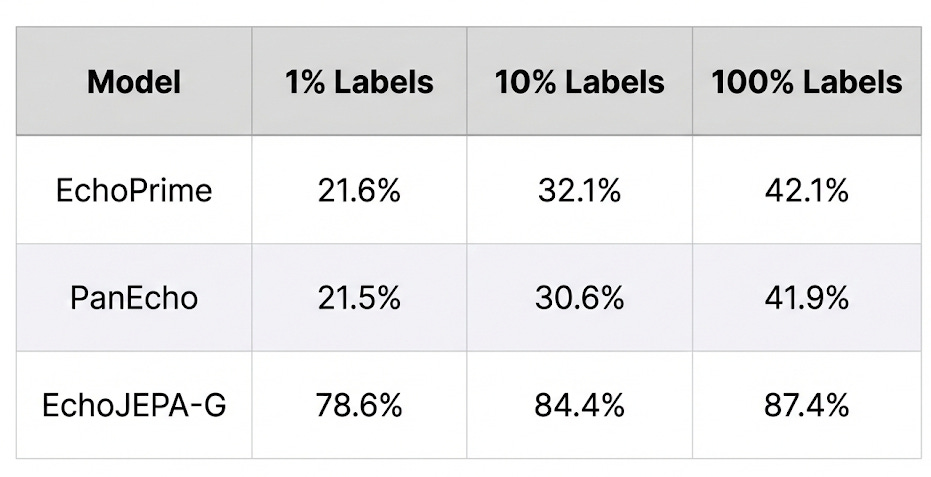
3. The 1% Labeled Data Result (This One Blew Our Minds)
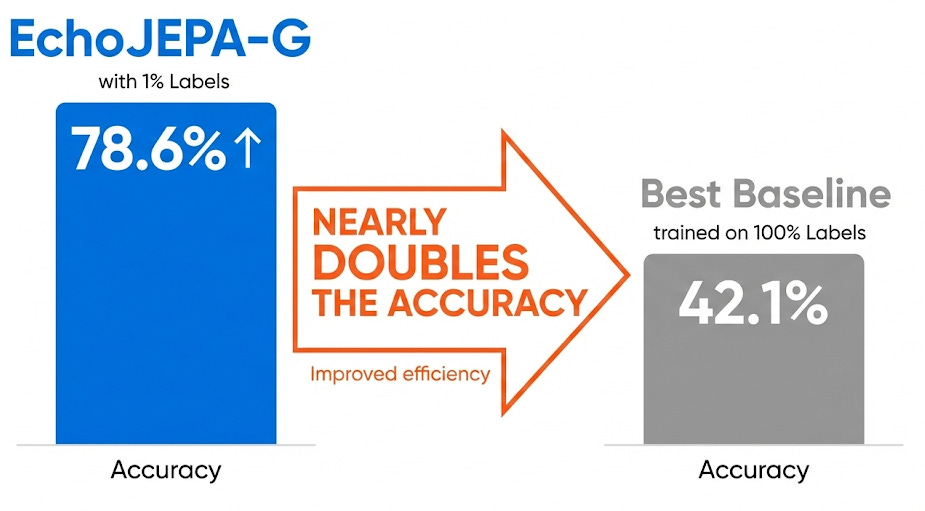
This is, to me, the most striking finding. How well can each model classify 12 standard echocardiographic views when given only 1% of labeled training data?
Read that again. EchoJEPA-G with 1% of labels (78.6%) nearly doubles the accuracy of the best baseline trained on 100% of labels (42.1%).
This means the model has already learned, from pure self-supervised observation, a rich internal representation of cardiac anatomy. The labeled data isn’t teaching it what the views are - it’s just giving names to concepts the model already understands.
Think about the clinical implications: hospitals with thousands of unlabeled echo studies sitting in archives could leverage those archives directly, needing only a tiny handful of annotations to build high-performing AI systems.
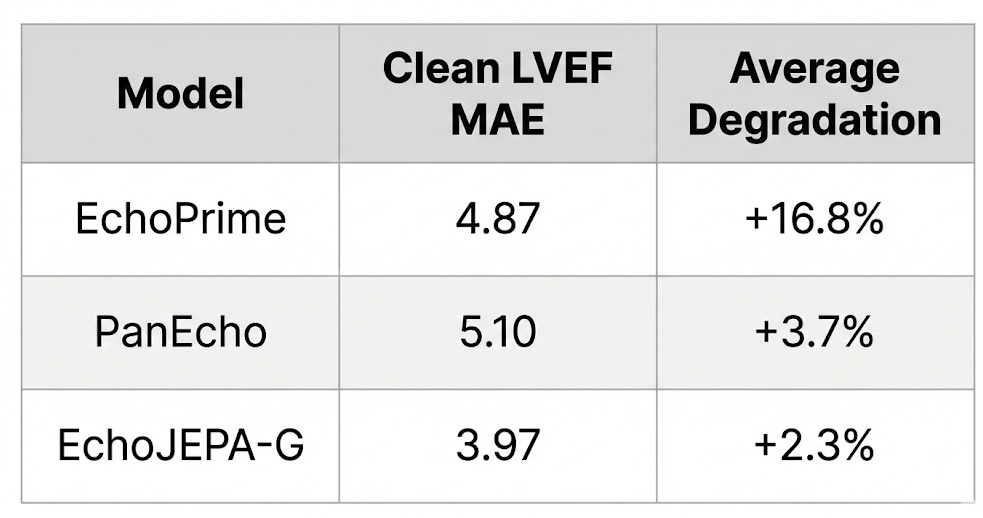
4. Robustness: When Image Quality Degrades
Here’s where it gets clinically important. Real-world echo quality varies enormously - patients with obesity have poor acoustic windows, ribs cast shadows, depth attenuates signal. The patients who need AI assistance the most are often the ones whose images are hardest to interpret.
We simulated these real-world degradations with physics-informed perturbations - depth attenuation (mimicking signal loss through tissue) and acoustic shadows (mimicking rib obstruction) - and measured how much each model’s performance deteriorated:
EchoJEPA is 86% less sensitive to acoustic artifacts than EchoPrime. Why? Because latent prediction never learned to care about speckle and acquisition noise in the first place. The model’s representations are anchored to stable anatomical structure, not surface-level pixel patterns.
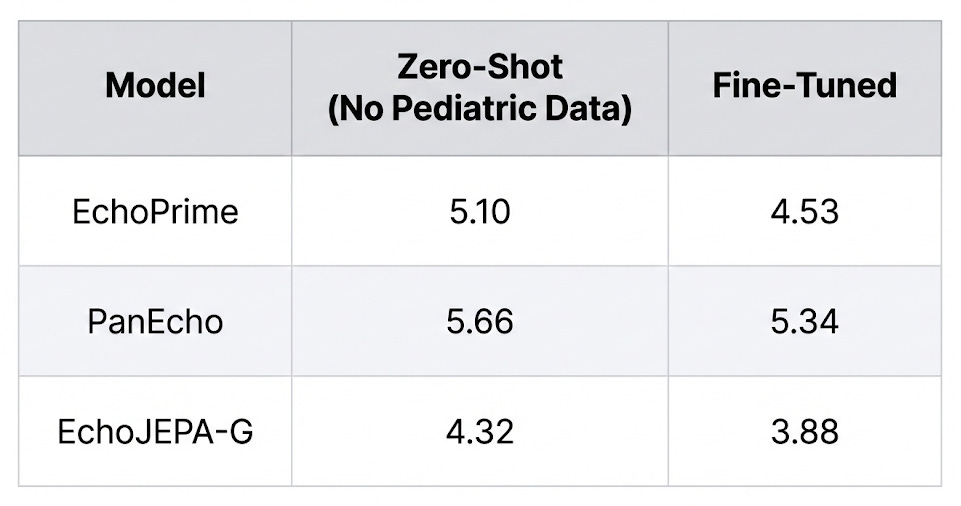
5. Zero-Shot Pediatric Transfer (Adults → Children, No Retraining)
Pediatric hearts are smaller, have different chamber proportions, and present different pathologies. We tested whether a model trained entirely on adult echo data could transfer to pediatric patients without any retraining:
EchoJEPA-G’s zero-shot performance on pediatric patients (4.32) beats every other model even after they’ve been fine-tuned on pediatric data. The model has never seen a child’s heart, and it’s already better than competitors that have.
This is what generalizable representations look like.
6. Multi-View RVSP Estimation
RVSP requires integrating information across multiple echocardiographic views - a clinically important and technically challenging task:
A 17% improvement over PanEcho on the Toronto benchmark, validating both the quality of the representations and the effectiveness of our multi-view fusion framework.
What the Model Actually “Sees”
One of the most compelling parts of the paper is the attention visualization. When you look at where each model focuses:
The model isn’t just predicting numbers. It’s interpreting the echocardiogram as a functional biological system. It understands the cardiac cycle.
The UMAP visualization of the frozen embedding space tells the same story: EchoJEPA forms distinct, well-separated clusters for different anatomical views — including a clean separation of transesophageal (TEE) from transthoracic (TTE) echocardiograms, without ever being told the difference. Baselines? Diffuse, overlapping blobs.
Why This Matters Beyond Benchmarks
Let me take off the researcher hat for a moment and put on the “person who’s worked in clinical AI at Philips and the University of Chicago” hat.
The bottleneck in medical AI has never been model architecture. It’s data annotation. Getting cardiologists to label 100,000 echo studies is prohibitively expensive and slow. Every hospital has massive archives of unlabeled echo data collecting digital dust.
EchoJEPA turns that liability into an asset. If a model can learn robust cardiac representations from unlabeled video alone - and then adapt to specific clinical tasks with minimal supervision - we fundamentally change the economics of medical AI development.
Here’s what this enables:
For resource-limited hospitals: Train high-performing AI with a fraction of the labeled data. A rural hospital with 50,000 unlabeled echoes and 500 labeled ones can now build systems that rival those from major academic centers.
For underserved patients: The patients who benefit most from automated echo analysis - those with obesity, lung disease, or limited acoustic windows - are exactly the ones whose images deviate most from training distributions. EchoJEPA’s robustness to image degradation means the model doesn’t fail on the patients who need it most.
For the research community: We are releasing EchoJEPA-L (trained on public MIMIC-IV-Echo data) and our entire evaluation framework. Frozen backbone + lightweight probe means clinical researchers can explore these representations without massive GPU clusters.
The Technical Contribution Summary
For my ML colleagues who want the quick reference:
What Comes Next
EchoJEPA opens several exciting directions: fine-grained tasks like valve segmentation, prospective validation on patients with poor acoustic windows, integration with interpretable reasoning frameworks for clinical decision support, and extension to other noisy modalities like fetal ultrasound and lung ultrasound.
The broader message is clear: the right self-supervised objective, matched to the physics of your domain, changes everything. Not all self-supervised learning is created equal. The choice of what you predict matters as much as how much data you train on.
Try It Yourself
EchoJEPA-L and our evaluation framework are open-source:
🔗 Paper: arxiv.org/abs/2602.02603 🔗 Code: github.com/alif-munim/EchoJEPA 🔗 Project Page: echojepa.com
This work is a collaboration between University Health Network, University of Toronto, Vector Institute, University of Chicago, UC San Francisco, Cohere Labs, and Philips Health. I’m grateful to co-lead this work alongside Alif Munim and Adibvafa Fallahpour, and to our incredible team of collaborators.
If you’re building AI for medical imaging - especially in noisy modalities like ultrasound - I’d love to hear from you. What’s the biggest bottleneck in your pipeline?
Comment. I read every response.
Teodora Szasz is a Senior Clinical Data Scientist at Philips and career coach who helps AI/ML professionals build careers they’re excited about. She writes Standout Systems, a daily newsletter on AI/ML and career development. Her work spans echocardiography, oncology imaging, and clinical AI deployment across regulated healthcare environments.