How I run clinical AI experiments in 2026: the full Claude Code system that produces FDA-ready evidence
The full system that takes evidence production from three weeks to three days: CLAUDE.md, three skills, four guardrail hooks, and the reproducible-research workflow. Plus the downloadable kit.
A note before you read
This post and the accompanying starter bundle are my personal work, developed independently of any employer or client.
The patterns, conventions, and code shared here are abstracted from general best practices in clinical AI engineering.
Not from any specific proprietary system, codebase, or workflow at my employer. Templates use intentionally generic examples and are not based on or derived from any specific real-world product.
Nothing here is endorsed by, attributable to, or disclosed by my employer or any client.Treat this as one data scientist’s personal framework.
Not as guidance from any organization.
If you adopt any of these patterns in a regulated environment, your own regulatory affairs, quality, and security teams must review them for fit with your specific context.
Three weeks ago, I produced the validation evidence package for a clinical AI model improvement in three days that would have taken me three weeks in 2024.
I did not ship the model. That is not my job.
As a senior clinical data scientist, my job is to design the study, run the experiments, and produce the evidence: the validation reports, the performance tables, the regulatory implication assessment.
That engineering, regulatory affairs, and clinical teams need to do their parts.
What changed is not that I ship faster. It is that the evidence I hand off is now traceable, reproducible, and submission-ready by default. Not a frantic week-of-deadline scramble to find the right config, the right git hash, the right validation numbers.
This is the post I wish I had read four years ago when I started doing clinical AI work in regulated environments.
It is the capstone of the hacker mindset series: the post where I show you exactly how the pieces fit together for a clinical data scientist working in a regulated environment, where the cost of sloppy traceability is measured in IRB findings and 510(k) deficiency letters.
This is post #10. (Full series: 1, 2, 3, 4, 5, 6, 7, 8, 9.)
What makes clinical AI different (the part everyone gets wrong)
Most data scientists learning AI tooling assume their workflow translates directly to clinical AI work.
It does not.
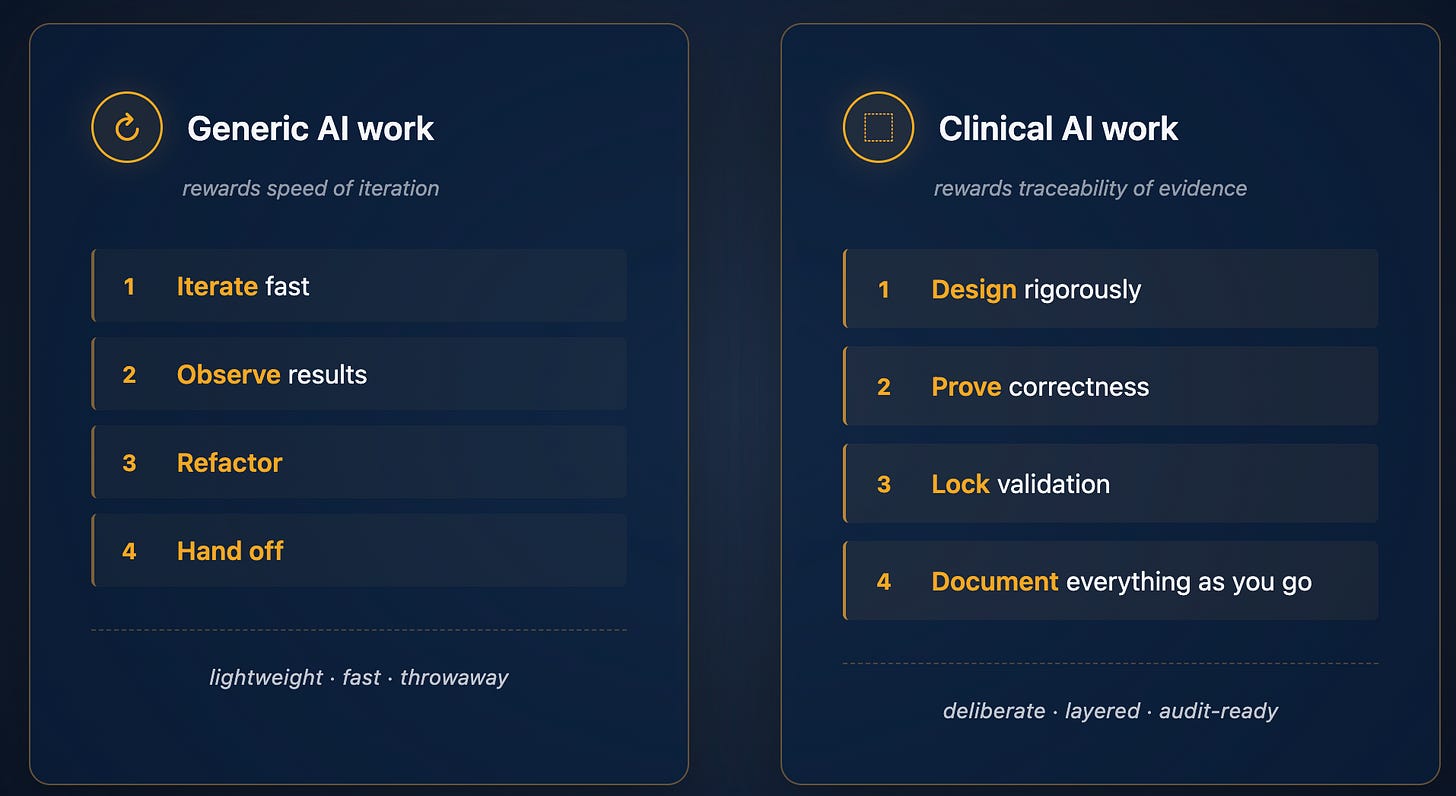
Generic AI work rewards speed of iteration.
Clinical AI work rewards traceability of evidence.
Every model output must be tied to the code that produced it, the data it was trained on, the validation slice that was held out, and the version of the pipeline that ran it. That is not bureaucratic overhead - it is what makes the evidence usable downstream by regulatory affairs, engineering, and clinical teams who have to act on it.
The mental model shift is the whole post:
Most data scientists who try to bring agentic AI tools into clinical work fail because they treat the agent like they would in a research notebook - ask for a quick result, accept the output, move on.
That approach falls apart the first time regulatory affairs asks “what data was this trained on?” and you cannot answer with confidence.
The system I am about to walk you through fixes that.
The agent moves at notebook speed; the documentation and traceability are produced automatically as a byproduct.
The clinical AI Claude Code stack - overview
Five layers, each building on the last:
I am going to walk you through the thinking behind each layer here, then give paid subscribers the full production-ready bundle below - every file, ready to drop into a clinical AI data science project this week.
Layer 1: The repo structure (the foundation)
This is where most clinical AI projects fail before they start.
The code, the experiments, the validation results, and the regulatory documentation all live in different places: different repos, different drives, different teams.
By the time you are six months into a project, nobody can answer “what version of the code produced figure 3 in the IRB submission?”
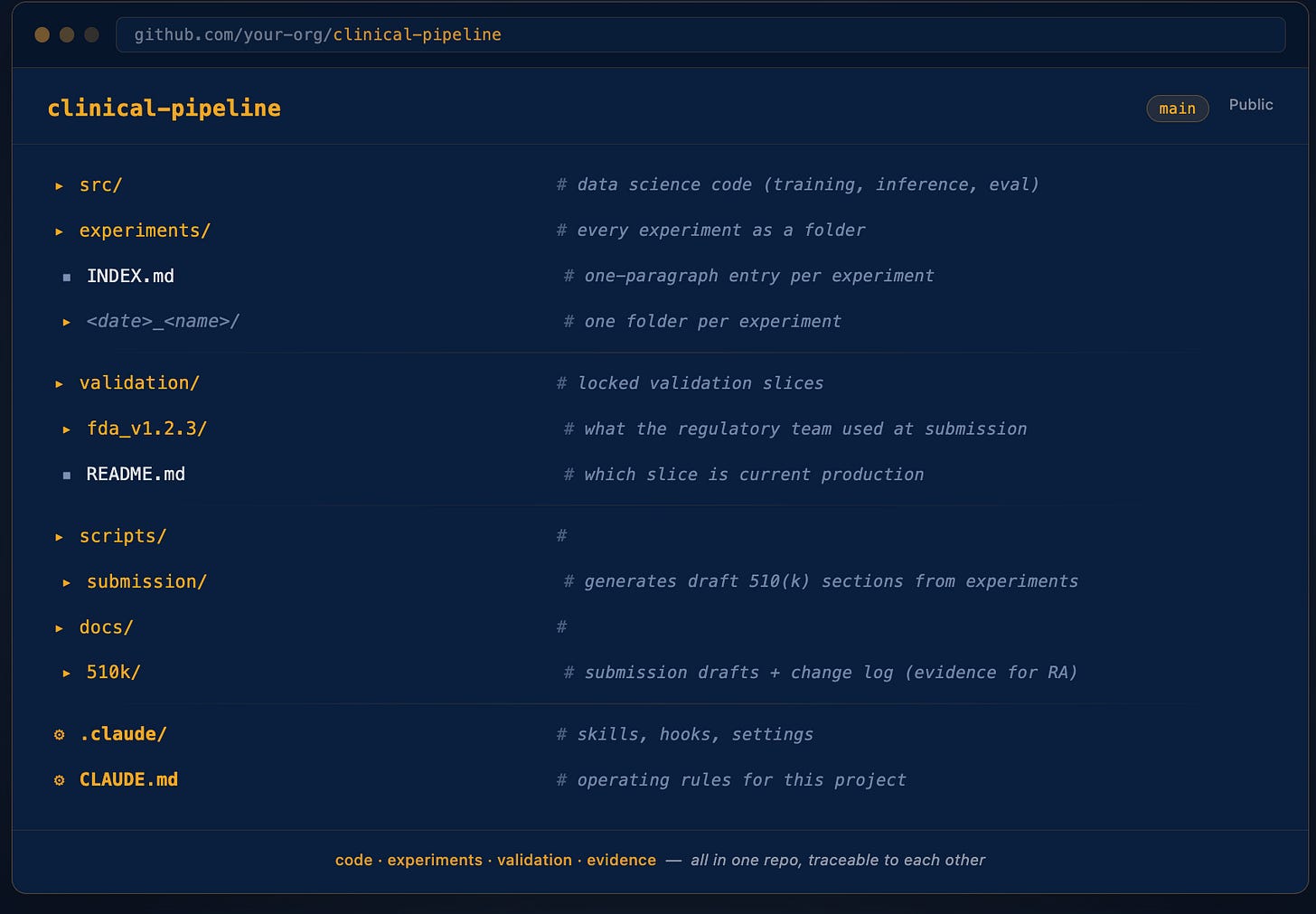
The fix is structural. Everything in one repo, organized so the relationship between code, experiment, result, and document is enforced by the directory layout itself.
A note on scope: this is the data science work area.
Production deployment lives in a separate repo owned by engineering.
The evidence handoff happens through docs/510k/drafts/ and experiments/INDEX.md, both of which downstream teams can read independently.
This separation of concerns is itself a regulatory hygiene best-practice - you do not want validation evidence and deployment configuration in the same place.
Three things matter about this layout:
1. Validation is a first-class directory, not an afterthought.
The validation/ folder contains frozen validation slices: the exact patient cohorts used in your FDA submission, IRB protocol, or most recent clinical validation report. These never change once locked.
They become the ground truth every future model change is measured against.
2. Experiments are reproducible artifacts, not throwaway notebooks.
Every experiment is its own folder.
The experiments/INDEX.md file is the single source of truth: one paragraph per experiment, grouped by phase.
When regulatory affairs asks “what experiments did you run for the LVEF feature,” you point them at INDEX.md and they can navigate to any run in three clicks.
3. Documentation is generated, not written.
The scripts/submission/ directory contains scripts that read your experiments and validation results and draft the relevant sections of submission documentation directly. No copy-paste of numbers from a notebook.
No “let me update figure 3 again.” If a number changes, you fix the experiment and rerun the script.
The draft updates automatically: and regulatory affairs, who actually owns the submission, gets a cleaner starting point.
That layout alone is worth a month of senior data scientist time on any clinical AI project. The bundle below contains the complete scaffolded version, ready to drop into a new project.
Layer 2: CLAUDE.md for clinical AI
You have seen the generic version in Post #6.
The clinical version adds five sections that change Claude Code’s behavior in the ways regulated environments require: Regulatory context, PHI rules, Validation rules, Experiment rules, Submission documentation rules.
Each of these does specific work.
The Regulatory context section tells Claude which model is locked and which validation slice is current: so Claude knows what it must not touch even when asked.
The PHI rules tell Claude that the scrubber lives at src/validators/phi.py::scrub() and that PHI is enforced by hooks, not by trust.
The Validation rules tell Claude that any change to inference-relevant code requires running the locked slice before the change can be considered evaluated and handed off.
The Experiment rules force every experiment through the /new-experiment skill so structure is consistent. The Submission rules establish that 510(k) section drafts are evidence packages for the regulatory team to review - never submitted directly from this repo.
Roughly 120 lines total. The bundle contains the full template with [EDIT] markers for everything project-specific. You spend twelve minutes filling in markers and your repo now has clinical-grade operating rules that every Claude Code session inherits automatically.
The single most valuable line in the whole file is the one establishing the production model path as immutable:
Production model path:
models/[version]/: NEVER MODIFY
That one line, combined with the validation lock hook (Layer 4), is what makes the difference between Claude Code as a clever toy and Claude Code as a tool you can responsibly use on clinical AI data science work that supports regulated software.
Layer 3: The three skills that compound
Skills (from Post #7) are reusable workflows captured as SKILL.md files.
Three of them are foundational for clinical AI data science:
/new-experiment: scaffolds a clinical experiment with config, pre-registered hypothesis, validation slice reference, and result template.
The hypothesis is the move.
Clinical AI experiments without a pre-registered hypothesis are not science: they are post-hoc rationalization. The skill refuses to proceed until you write one.
/run-validation: executes the locked validation slice and reports regression vs. baseline.
The critical step: it verifies the slice’s manifest hash before running.
If the validation files have been silently modified since the last submission, the skill refuses to proceed and flags the integrity issue.
Validation slices that can be silently modified are not locked validations.
This is the kind of small thing that takes you from “I think this is reproducible” to “I can prove this is reproducible to a regulatory auditor.”
/draft-510k-section: reads the change log and the validating experiments linked from it, then drafts the submission section in regulatory-affairs-ready format.
It always ends with a non-optional reminder: this is a draft, regulatory affairs review is required before submission, the data scientist’s job ends at producing the evidence package. Make that clear every single time, or someone will eventually treat the draft as final.
Each skill is a small markdown file - 20-30 lines.
The bundle contains all three, ready to drop into .claude/skills/.
Layer 4: The hooks that make it real
Hooks (from Post #8) are deterministic guardrails that fire whether the agent wants them to or not. For clinical AI work, four hooks form the core safety system.
phi_guard.sh (PreToolUse): blocks any write containing PHI patterns.
Catches SSN, MRN, DICOM patient names (the LASTNAME^FIRSTNAME caret format), and patient DOB near patient-name keywords.
Allows writes to data/raw/ (where raw PHI legitimately lives, but which is locked by the next hook).
validation_lock.sh (PreToolUse): refuses any modification to validation/fda_*/, models/v*/, or data/raw/.
Catches both direct file writes and shell commands targeting those paths (no sneaking past it with rm -rf or output redirection).
audit_trail.sh (PostToolUse): logs every file modification to an append-only audit log with timestamp, user, file path, and git hash at the time of the change. Combined with git history, you get a complete provenance chain that any regulatory auditor can follow.
change_log_check.sh (PostToolUse): warns (does not block) when src/pipeline/ is modified without a corresponding docs/510k/change_log.md entry.
The blocking happens at PR time via CI, not at session time - because in-development work legitimately modifies pipeline code many times before the change log entry is written.
The hooks are roughly 30-80 lines each.
The bundle contains all four, the .claude/settings.json that wires them together, and a tests/test_hooks.sh script that adversarially tests them (16 cases, all passing).
A safety hook without a test is theater.
The test file is what makes the hooks defensible in a regulatory review.
And the part that makes the entire system actually work in practice: the CI integration.
Local hooks can be bypassed (you can always edit them or comment them out). The GitHub Actions workflow runs the same checks at PR time, where they cannot be bypassed without an explicit override visible to your team.
The bundle includes .github/workflows/clinical-checks.yml with three CI jobs - PHI scan on the PR diff, validation lock check, and changelog-required check.
Without the CI integration, the hooks are theater. With it, they are infrastructure that the regulatory team can trust.
Layer 5: The reproducible-research workflow
The index-and-detail pattern from the workshop deck I built this series on, fully adapted for clinical AI data science.
The principle: every experiment is a self-contained artifact. The experiments/INDEX.md file gives you a one-paragraph view of every experiment ever run on the project.
Each entry links to a detail page in experiments/<date>_<name>/NOTES.md that contains everything needed to reproduce or interpret the run: hypothesis, exact command, git hash of the code at run time, config path with SHA-256, dataset version, validation slice used, observed numbers, plots, and the field that nobody else writes —
Regulatory Implication.
This is the single most valuable field in any clinical AI experiment note.
It forces the data scientist to do an initial assessment:
Does this change appear to warrant a 510(k) supplement? An IRB amendment? A clinical re-validation? Does it appear to affect labeling or intended use? Does it appear to introduce new failure modes?
If the answer to all of these is “no impact,” state that explicitly with reasoning.
Empty here means forgotten, not no impact.
Critically, this is data-scientist input to a regulatory-affairs decision - not the final regulatory determination.
Your job is to provide a clear, structured starting point. Regulatory affairs makes the actual call.
The other discipline that makes the whole system regulatory-grade: fix the script, not the output. If a number in your draft submission section is wrong, you do not edit the figure or the table.
You fix the underlying data or the script that generated it, and you rerun. The draft updates automatically.
Every number you hand to regulatory affairs is traceable to an experiment, which is traceable to a config, which is traceable to a git hash, which is traceable to a code change.
The full chain is auditable, by construction.
The bundle contains the templates for INDEX.md and NOTES.md, plus the convention for artifact-generation scripts.
What this looks like in practice (the 3-day evidence package)
Three weeks ago, I produced an evidence package that would have taken three weeks of work in 2024. The condensed version:
Day 1, morning. I opened Claude Code in the project and typed three sentences: “Here is a model improvement engineering wants me to evaluate. Before you start, ask me 5 questions to clarify the study design and constraints. Then propose the verification clause for the validation.”
Claude asked the questions. The conversation became the study design. The verification clause we settled on was four lines long: the locked FDA validation slice must execute cleanly, the regression thresholds must all pass, the experiment must register in the index, and the regulatory implication assessment must be written before the experiment closes.
Day 1, afternoon. I ran /new-experiment to scaffold the study. I wrote the hypothesis. Claude generated the evaluation harness while the hooks watched for PHI leaks and validation-lock violations in the background. The first run completed; one metric came in within regression threshold but suspicious enough to warrant a second look. Caught by the verification clause forcing me to write the regulatory implication field - which is what made me stop and investigate before submitting flawed evidence downstream.
Day 2. I re-ran the validation after fixing the metric drift, all numbers passed, and I generated the canonical figures via the artifact scripts. I wrote a one-paragraph entry in docs/510k/change_log.md linking back to the experiment.
Day 3. I ran /draft-510k-section "Performance Testing". Claude read the change log, followed the link to the experiment notes, generated the section draft with the validation numbers in the canonical table format, and put the draft in docs/510k/drafts/. I reviewed the draft once and handed it to regulatory affairs. They had two small edits - both stylistic, not substantive. The numbers, the figures, the references, the regulatory implication assessment were all correct because they were generated from the same source of truth that ran the validation.
Three days. Zero scrambling. Zero “wait, which version of the model did we run on which slice?”
The traceability was built in because the system enforced it, not because I remembered to be disciplined.
The evidence package made it through regulatory review smoothly. Engineering reviewed the underlying code change and merged it cleanly. Clinical signed off on the implication assessment with one clarifying question. All three teams downstream of me were unblocked by the quality of the package - not by my involvement in their work.
That is the workflow the bundle below sets up.
What’s in the bundle (for paid subscribers)
The full production-ready starter, downloadable below.
Drop it into any clinical AI data science project, configure the project-specific paths, and you have a clinical-AI-aware Claude Code setup running in twenty minutes.
38 files. Every shell hook verified against an adversarial test suite.
Every Python file syntax-checked.
Every safety guarantee tested.
The bundle is for the data scientist’s part of clinical AI work: experiment discipline, validation rigor, evidence production, draft documentation generation.
It explicitly does not include things owned by other teams: deployment infrastructure, QMS artifacts, regulatory submission filing tools, clinical study management, or security tooling. Those belong to engineering, QA, regulatory affairs, clinical operations, and security respectively.
The bundle is licensed permissively (MIT) so you can adopt any portion of it for your own work.
The included LICENSE and NOTICE files make explicit that this is educational scaffolding, not validated software - your own regulatory affairs, quality, and security teams must review it for fit with your specific context before any production use.